|
About Unsplicer
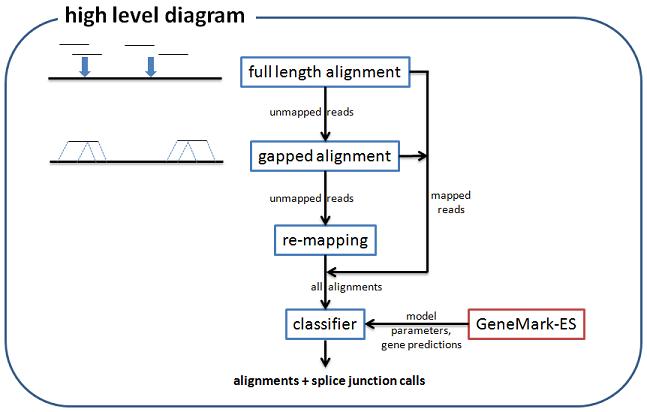
UnSplicer is an RNA-seq alignment program, providing fast and accurate alignment of short reads
to a reference genome. UnSplicer is different from most alignment programs in that it uses DNA
sequence models to improve detection of splice junctions.
UnSplicer requires two inputs which are provided by the output of GeneMark-ES: HMM model
parameters and ab initio gene predictions. In order to run UnSplicer, you should
have successfully run GeneMark-ES for your genome.
UnSplicer is a sister pipeline to TrueSight.
UnSplicer and TrueSight each employ a method for classifying splice junctions as either "true" or "false."
There are three major differences between UnSplicer and TrueSight:
UnSplicer and TrueSight share many components, including the pipeline for aligning fully exonic reads
(no gaps), and the program which implements the "anchor and extend" strategy for initial gapped alignment.
More details can be found in the UnSplicer publication.
Mark Borodovsky
|
|